—Accelerating the rational design of catalytic systems by combining machine learning and quantum chemical calculations—
Key Points
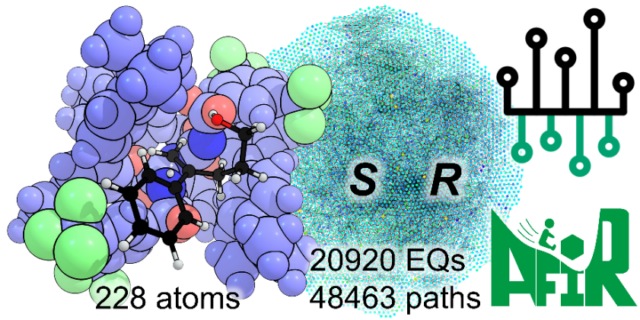
- Successfully constructed a reaction path network for a large catalytic system comprising over 200 atoms.
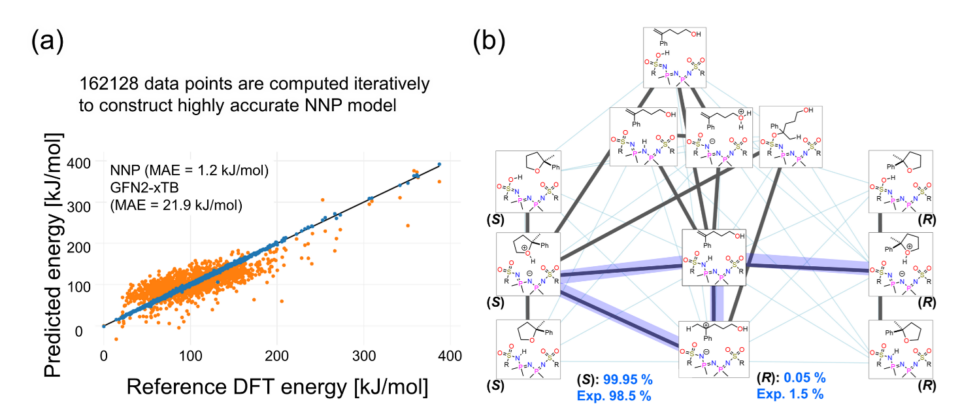
- Theoretically reproduced the experimental enantioselectivity through kinetic analysis utilizing approximately 46,000 reaction pathways.
- Based on the NNP/AFIR method, a catalyst mechanism analysis method has been established which is not dependent on prior assumptions.
Overview
A research group led by Professor Satoshi Maeda, WPI-ICReDD and the Graduate School of Science at Hokkaido University, has successfully constructed a large-scale reaction path network of an asymmetric catalytic system using a new computational method that combines machine learning and quantum chemical calculations, and has theoretically reproduced the high experimental enantioselectivity.
Although asymmetric catalysts are essential for the synthesis of pharmaceuticals and functional materials, their large size and highly flexible geometries, making it difficult to theoretically understand how stereoselectivity arises. In this study, the researchers employed the “NNP/AFIR method”—a combination of the neural network potential (NNP) and the AFIR (Artificial Force Induced Reaction) method—to a catalytic system comprising over 200 atoms. As a result, they constructed a large-scale reaction path network consisting of 20,920 equilibrium structures and 48,463 reaction pathways.
Furthermore, the rate constant matrix contraction (RCMC) method, a kinetic simulation method based on the obtained reaction path network, explained the high experimental enantioselectivity. This study demonstrates a new theoretical framework capable of analyzing complex asymmetric catalytic systems without prior assumptions.
The results of this study were published in ACS Central Science on Monday, March 30, 2026 (Japan Standard Time).